Getting started¶
You can use DASH3R through the graphical user interface (GUI) or command line:
For GUI¶
To start the GUI, type
dash3r
A GUI that looks like the image below should appear. You can hover any of buttons in the GUI to see a brief description of the command.
 Graphical user interface for the Dasher application
Graphical user interface for the Dasher application
You can get the command usage info by click the “Help” box on any of the pop-up windows.
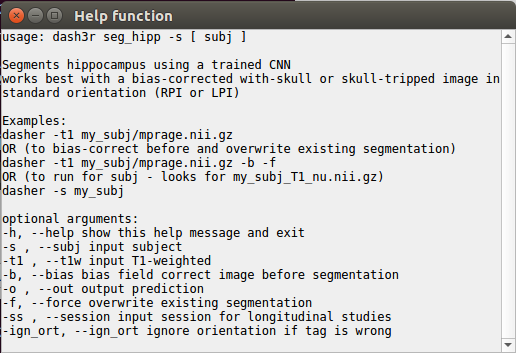 Help screen for graphical user interface for Dasher application
Help screen for graphical user interface for Dasher application
For Command Line¶
You can see all the dasher commands by typing either of the following lines:
dash3r -h
dash3r --help
Once you know the command you want to know from the list, you can see more information about the command. For example, to learn more about seg_hfb:
dash3r seg_hipp -h
dash3r seg_hipp --help
Hippocampal volumes¶
To extract hippocampal volumes use the GUI (Stats/Hippocampal Volumes) or command line:
dash3r stats_hp -h
QC¶
QC files are automatically generated in a sub-folder within the subject folder. They are .png images that show a series of slices in the brain to help you quickly evaluate if your command worked successfully, especially if you have run multiple subjects. They can also be created through the GUI or command line:
dash3r seg_qc -h
The QC image should look like this:
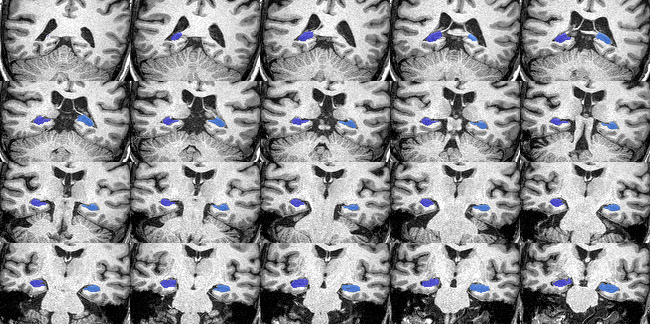 Quality control imagefor hippocampus segmentation
Quality control imagefor hippocampus segmentation
Logs¶
Log files are automatically generated in a sub-folder within the subject folder. They are .txt files that contain information regarding the command and can be useful if something did not work successfully.
File conversion¶
Convert Analyze to Nifti (or vice versa)
dash3r filetype
Required arguments:
-i , --in_img input image, ex:MM.img
-o , --out_img output image, ex:MM.nii
Example:
dash3r filetype --in_img subject_T1.img --out_img subject_T1.nii.gz